The authors looked at how soil temperature changes with fire to develop a sensor system that could aid in earlier detection of fires.
Read More...Browse Articles
Predicting and explaining illicit financial flows in developing countries: A machine learning approach
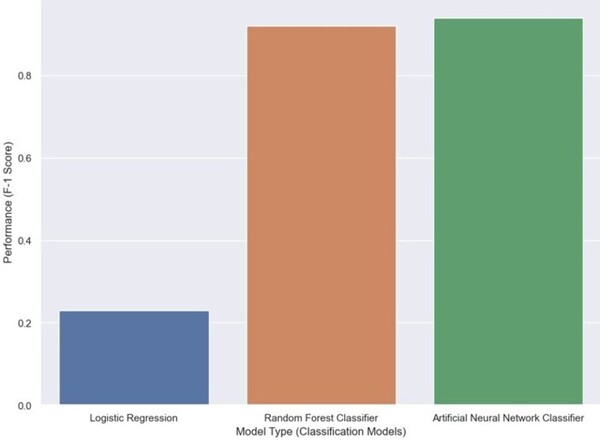
The authors looked at the ability of different machine learning algorithms to predict the level of financial corruption in different countries.
Read More...How artificial intelligence deep learning models can be used to accurately determine lung cancers
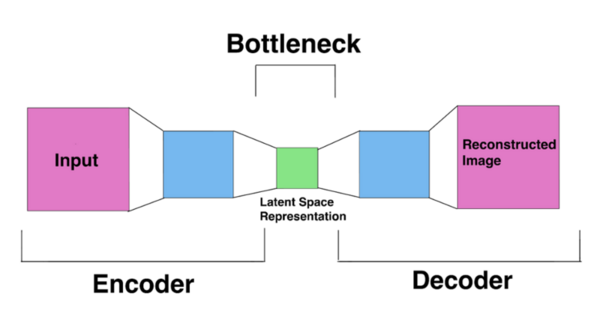
The authors looked at the ability of different deep learning models to predict the presence of lung cancer from chest CT scans. They found that a pre-trained CNN model performed better than an autoencoder model.
Read More...Levering machine learning to distinguish between optimal and suboptimal basketball shooting forms
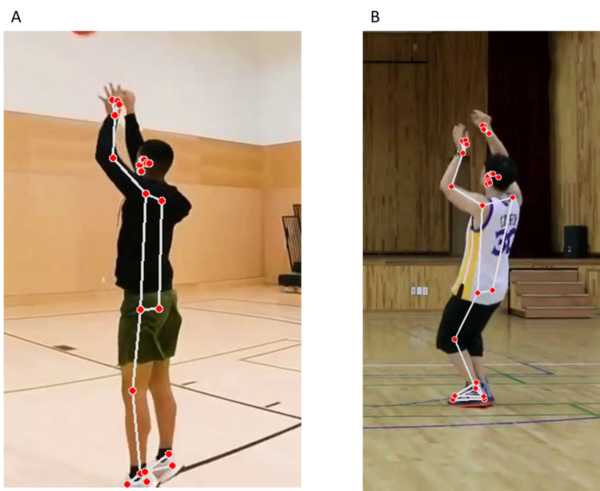

The authors looked at different ways to build computational resources that would analyze shooting form for basketball players.
Read More...Comparing transformer and RNN models in BCIs for handwritten text decoding via neural signals
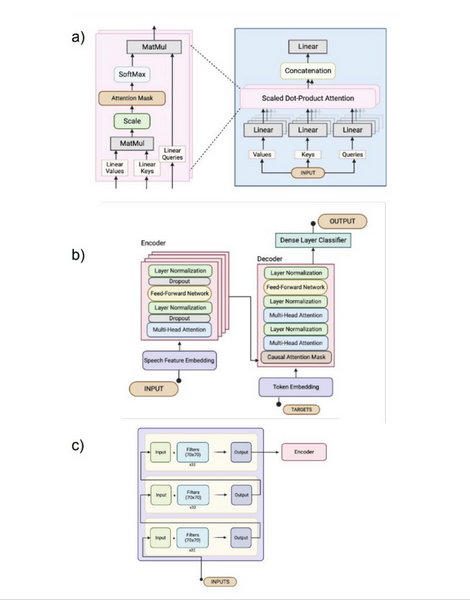
Brain-Computer Interface (BCI) allows users, especially those with paralysis, to control devices through brain activity. This study explored using a custom transformer model to decode neural signals into handwritten text for individuals with limited motor skills, comparing its performance to a traditional RNN-based BCI.
Read More...Unlocking robotic potential through modern organ segmentation
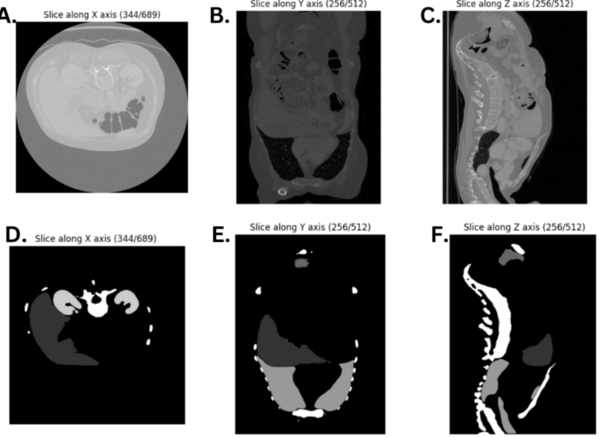
The authors looked at different models of semantic segmentation to determine which may be best used in the future for segmentation of CT scans to help diagnose certain conditions.
Read More...Large Language Models are Good Translators
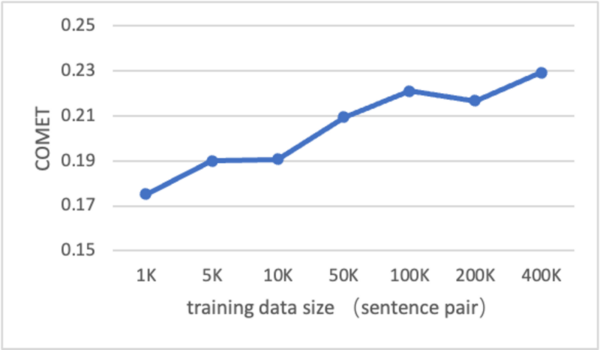
Machine translation remains a challenging area in artificial intelligence, with neural machine translation (NMT) making significant strides over the past decade but still facing hurdles, particularly in translation quality due to the reliance on expensive bilingual training data. This study explores whether large language models (LLMs), like GPT-4, can be effectively adapted for translation tasks and outperform traditional NMT systems.
Read More...Identifying shark species using an AlexNet CNN model
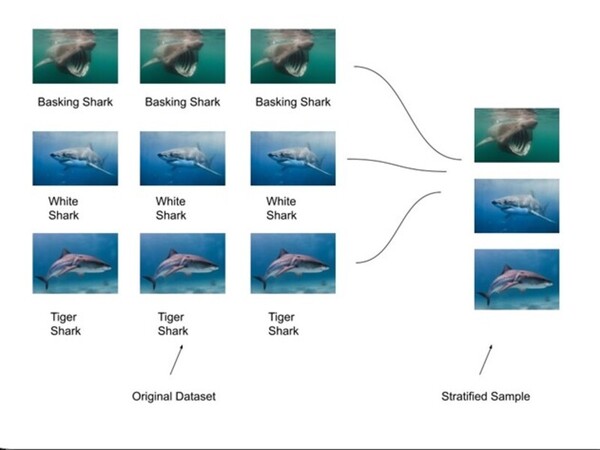
The challenge of accurately identifying shark species is crucial for biodiversity monitoring but is often hindered by time-consuming and labor-intensive manual methods. To address this, SharkNet, a CNN model based on AlexNet, achieved 93% accuracy in classifying shark species using a limited dataset of 1,400 images across 14 species. SharkNet offers a more efficient and reliable solution for marine biologists and conservationists in species identification and environmental monitoring.
Read More...Transcriptomic profiling identifies differential gene expression associated with childhood abuse
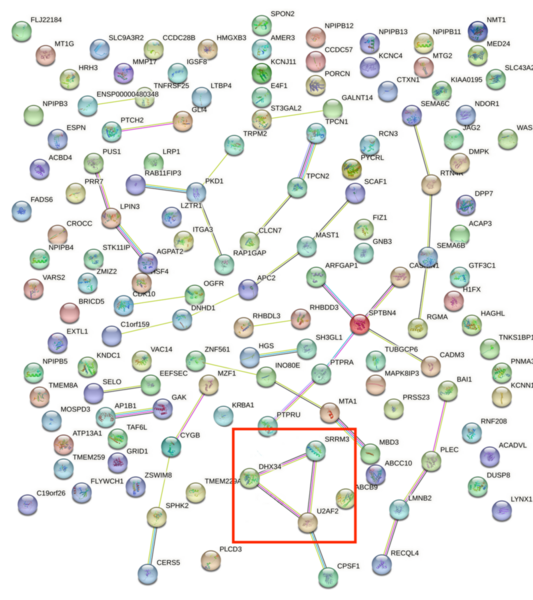
Childhood abuse has severe and lasting effects throughout an individual's life, and may even have long-term biological effects on individuals who suffer it. To learn more about the effects of abuse in childhood, Li and Yearwood analyze gene expression data to look for genes differentially expressed genes in individuals with a history of childhood abuse.
Read More...Predictions of neural control deficits in elders with subjective memory complaints and Alzheimer’s disease
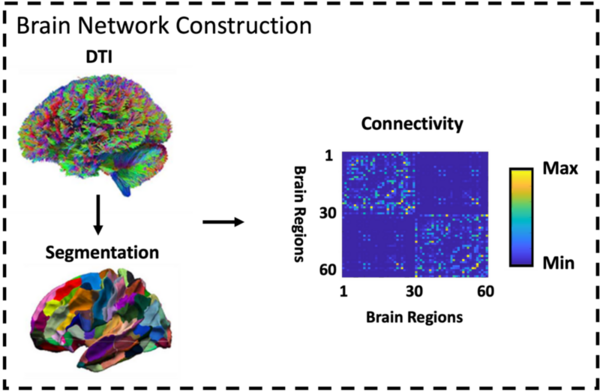
The authors compare neuroimaging datasets to identify potential new biomarkers for earlier detection of Alzheimer's disease.
Read More...