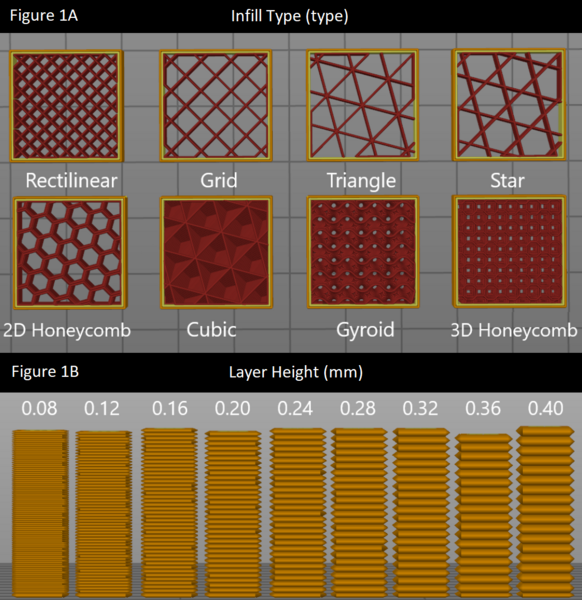
In this study, the authors test different infill patterns to determine which would be the strongest and most durable for 3D printing applications, which have become an integral part of many facets of life.
Read More...Optimizing 3D printing parameters: Evaluating infill type and layer height effects on tensile fracture force

In this study, the authors test different infill patterns to determine which would be the strongest and most durable for 3D printing applications, which have become an integral part of many facets of life.
Read More...Awareness of plastic pollution and adoption of green consumer lifestyles among students from high school

In this study, the authors test ways to increase knowledge of green consumerism amongst high school students. Their knowledge was measured based on the New Ecological Paradigm Scale.
Read More...Effect of mass and center of gravity on vehicle speed and braking performance

In this study, the authors test whether a gravity vehicle, which is a vehicle powered by its own gravity on a ramp, could be designed to move faster when mathematical calculations for optimal mass and center of gravity were applied in the design.
Read More...Assessing and Improving Machine Learning Model Predictions of Polymer Glass Transition Temperatures
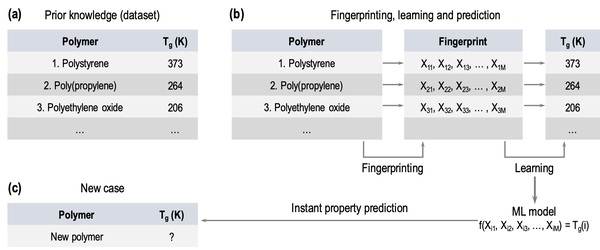
In this study, the authors test whether providing a larger dataset of glass transition temperatures (Tg) to train the machine-learning platform Polymer Genome would improve its accuracy. Polymer Genome is a machine learning based data-driven informatics platform for polymer property prediction and Tg is one property needed to design new polymers in silico. They found that training the model with their larger, curated dataset improved the algorithm's Tg, providing valuable improvements to this useful platform.
Read More...Modeling Energy Produced by Solar Panels
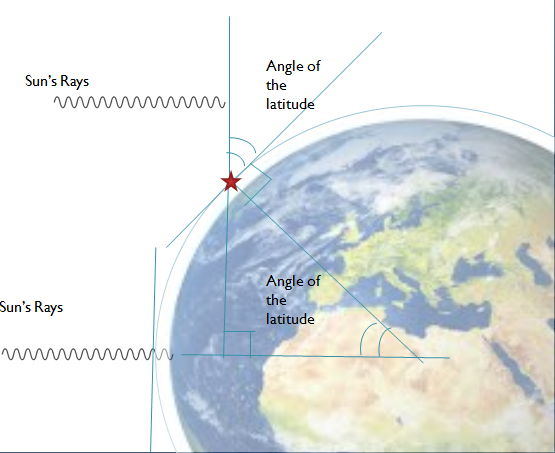
In this study, the authors test the effect that the tilt angle of a solar panel has on the amount of energy it generates. This investigation highlights a simple way that people can harvest renewable energy more efficiently and effectively.
Read More...The Effects of Ocean Acidification on the food location behavior and Locomotion of Pagurus Longicarpus
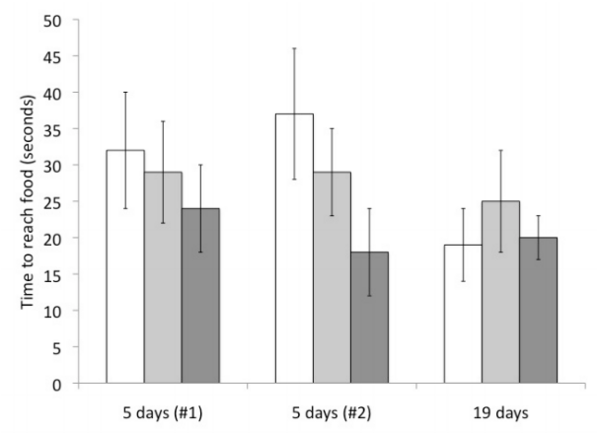
Increasing levels of atmospheric carbon dioxide is slowly acidifying our oceans. Here the authors test the effects of ocean acidification on the ability of hermit crabs (P. longicarpus) to find food. Though no statistically significant changes in food finding were observed, the data suggest a trend toward different activity.
Read More...De novo design of a dual-target inhibitor against tau phosphorylation and acetylation for Alzheimer's therapy
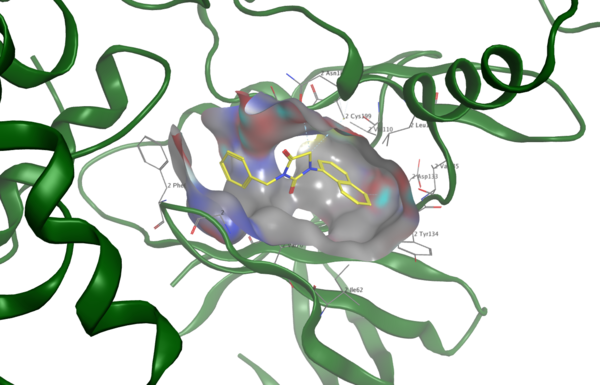
The authors use computational methods to compare tau acetylation to the better studied tau phosphorylation in Alzheimer's disease and then design and computationally test a new drug to prevent abnormal post-translational modifications of tau.
Read More...Effects of common supplements on human platelet aggregation in vitro
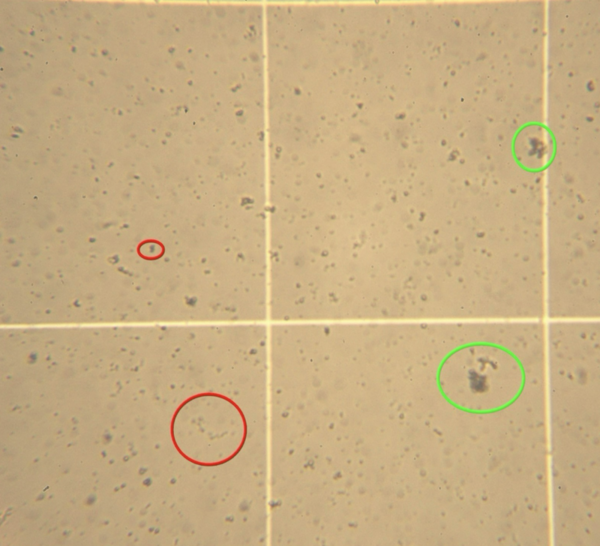
There is a need for safe and effective therapies to prevent platelet aggregation associated with cardiovascular diseases. Prabhakar and Prabhakar test to see whether dietary supplements claiming to reduce cardiovascular disease risk will affect aggregation of human platelets.
Read More...Assessing the possibility of using entomopathogenic fungi for mosquito control in Hawaii
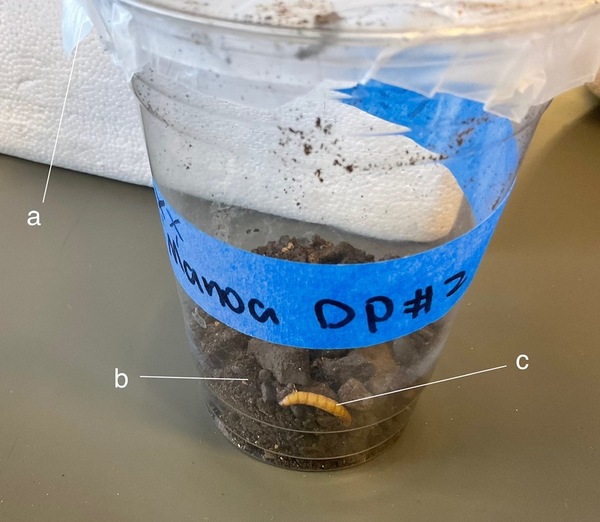

Fungi that attack and kill insects have promise for targeting mosquitoes without the harmful environmental impacts of chemicals like DDT. To find out whether fungi might be effective in controlling mosquitoes in Hawaii, Jiang and Chan test the effects of Hawaiian fungal isolates on mosquito larvae.
Read More...Predicting smoking status based on RNA sequencing data
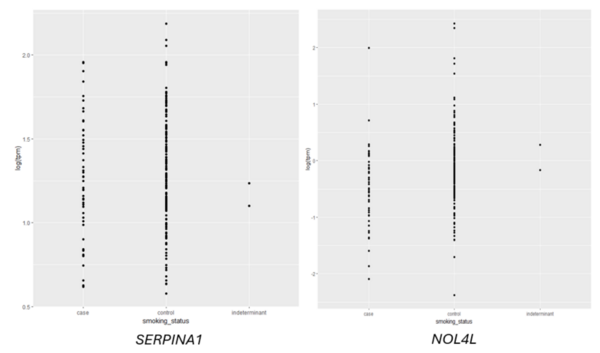
Given an association between nicotine addiction and gene expression, we hypothesized that expression of genes commonly associated with smoking status would have variable expression between smokers and non-smokers. To test whether gene expression varies between smokers and non-smokers, we analyzed two publicly-available datasets that profiled RNA gene expression from brain (nucleus accumbens) and lung tissue taken from patients identified as smokers or non-smokers. We discovered statistically significant differences in expression of dozens of genes between smokers and non-smokers. To test whether gene expression can be used to predict whether a patient is a smoker or non-smoker, we used gene expression as the training data for a logistic regression or random forest classification model. The random forest classifier trained on lung tissue data showed the most robust results, with area under curve (AUC) values consistently between 0.82 and 0.93. Both models trained on nucleus accumbens data had poorer performance, with AUC values consistently between 0.65 and 0.7 when using random forest. These results suggest gene expression can be used to predict smoking status using traditional machine learning models. Additionally, based on our random forest model, we proposed KCNJ3 and TXLNGY as two candidate markers of smoking status. These findings, coupled with other genes identified in this study, present promising avenues for advancing applications related to the genetic foundation of smoking-related characteristics.
Read More...