An Analysis of Soil Microhabitats in Revolutionary War, Civil War, and Modern Graveyards on Long Island, NY
(1) William Floyd High School, Long Island, New York, (2) William Floyd High School, Long Island, NY, (3) Cold Spring Harbor Laboratory, Long Island, New York
https://doi.org/10.59720/18-017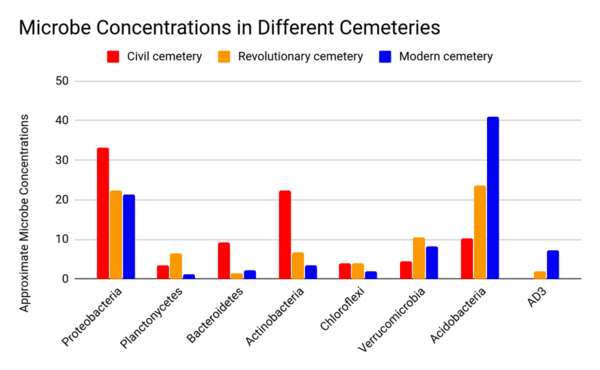
Previously established data indicate that cemeteries have contributed to groundwater and soil pollution. This study aimed to determine whether there was a variation in the microbes in cemeteries due to embalming techniques and to identify primary microbial communities in each cemetery. Different embalming techniques were used during different eras in recent history. These fluids can impact both epinecrotic and thanatomicrobiomes, the microbiomes that exist in decomposing remains. Additionally, these fluids can seep into the microbial communities, affecting the microbiome composition, and continue to leach into the larger surrounding environment, contributing to water source and aquifer pollution. We evaluated the 16S rDNA microbial gene followed by QIIME analyses using the Jupyter Notebooks application. We hypothesized that microbial variation would be high between cemeteries of different eras due to dissimilarities between embalming techniques employed, and furthermore, that specific microbes would act as an indication for certain contaminants. The results indicate that the following phyla were present: Proteobacteria, Planctomycetes, Bacteroidetes, Actinobacteria, Chloroflexi, Verrucomicrobia, Acidobacteria, and AD3. These taxonomic classifications were present in all sites with the exception of AD3 which was absent in the Civil War Cemetery. Overall, cemetery sites were clustered together based on location due to variations in the concentrations of the phyla and their more specific taxa.
This article has been tagged with: